Informácie o bakalárskej práci
Meno študenta: Sára Guziová
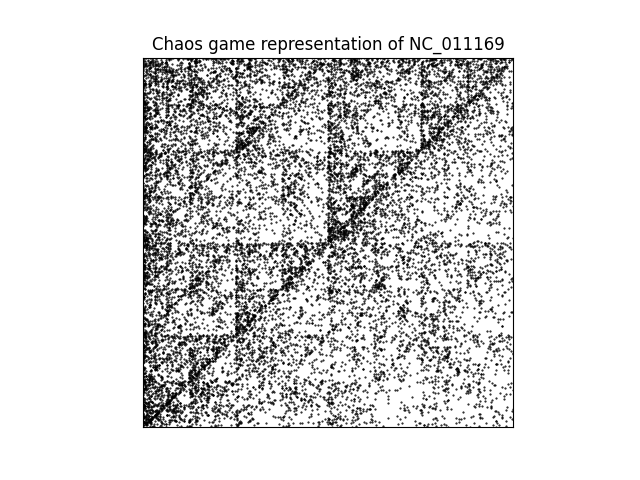
Názov práce: Teória hry chaosu a jej použitie na porovnanie postupností v bioinformatike
Meno školiteľa: prof. RNDr. Mária Lucká, PhD.
Kontakt na študenta: guziova6@uniba.sk
Zadanie
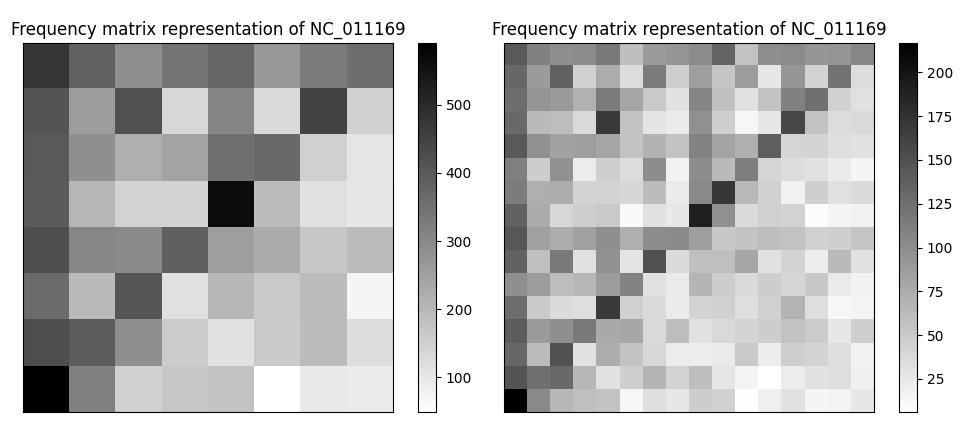
Výpočet podobnosti medzi dvomi nukleotidovými sekvenciami je jedným zo základných problémov bioinformatiky. Súčasné metódy sú založené buď na výpočtovo náročnom zarovnaní sekvencií alebo na použití metód bez zarovnania. Nové možnosti v tomto smere prináša reprezentácia postupností pomocou hry chaosu, ktorá využíva grafické prostredie a transformuje postupnosti rôznej dĺžky na obrazy alebo matice rovnakej veľkosti. Takáto reprezentácia je vhodná aj na kódovanie čŕt v strojovom učení.
Cieľom bakalárskej práce je aplikovať teóriu hry chaosu na hľadanie podobnosti veľkých genomických postupností a porovnať ju z hľadiska presnosti s inými metódami bez zarovnania na vybraných dátových množinách.
Zoznam zdrojov
- Jonas S Almeida. Sequence analysis by iterated maps, a review. Briefings in
bioinformatics, 15(3):369–375, 2014.
- Gaëtan Benoit, Claire Lemaitre, Dominique Lavenier, Erwan Drezen, Thibault
Dayris, Raluca Uricaru, and Guillaume Rizk. Reference-free compression of high
throughput sequencing data with a probabilistic de bruijn graph. BMC bioinfor
matics, 16(1):1–14, 2015.
- Broňa Brejová and Tomáš Vinař. Metódy v bioinformatike. Fakulta matematiky,
fyziky a informatiky Univerzita Komenského v Bratislave, 2011.
- Madison Cohen-McFarlane, Kevin Dick, James R Green, and Rafik Goubran.
Chaos game representation of audio signals. In 2021 IEEE International Instrumentation and Measurement Technology Conference (I2MTC), pages 1–6. IEEE,
2021.
- Megan C. Conroy et al. Uk biobank: a globally important resource for cancer research. British Journal of Cancer, 128(4):519–527, 2023
- Constantin P Cristescu, Cristina Stan, and Eugen I Scarlat. Modeling with the
chaos game (i). simulating some features of real time series. UPB Sci Bull Ser A,
71:95–100, 2009.
- Fatima Cvrčková. Úvod do praktické bioinformatiky. Academia, 2006.
- Nicki Skafte Detlefsen, Søren Hauberg, and Wouter Boomsma. Learning meaningful representations of protein sequences. Nature communications, 13(1):1914,2022
- Tomáš Farkaš, Jozef Sitarčík, Broňa Brejová, and Mária Lucká. Swspm: A novel
alignment-free dna comparison method based on signal processing approaches.
Evolutionary Bioinformatics, 15:1176934319849071, 2019.
- Umesh Ghoshdastider and Banani Saha. GenomeCompress: a novel algorithm for
DNA compression, 2005.
- Rosario Gilmary, Akila Venkatesan, and Govindasamy Vaiyapuri. Compression
techniques for DNA sequences: A thematic review. J. Comput. Sci. Eng., 15(2):59–
71, 2021.
- H Joel Jeffrey. Chaos game representation of gene structure. Nucleic acids research, 18(8):2163–2170, 1990.
31
- Jijoy Joseph and Roschen Sasikumar. Chaos game representation for comparison
of whole genomes. BMC bioinformatics, 7(1):1–10, 2006.
- Arthur M. Lesk. bioinformatics. [Citované 2024-01-10] Dostupné na https://www.britannica.com/science/bioinformatics.
- LibreTexts. Storing genetic information. [Citované 2023-05-20] Dostupné
na https://bio.libretexts.org/Courses/Lumen_Learning/Biology_for_Non-Majors_I_%28Lumen%29/08%3A_DNA_Structure_and_Replication/8.02%3A_Storing_Genetic_Information.
- Hannah Franziska Löchel and Dominik Heider. Chaos game representation and its
applications in bioinformatics. Computational and structural biotechnology journal, 19:6263–6271, 2021.
- Vijini Mallawaarachchi. Pairwise sequence alignment using biopython.
[Citované 2024-01-25] Dostupné na https://towardsdatascience.com/
pairwise-sequence-alignment-using-biopython-d1a9d0ba861f.
- Knižnica matplotlib. [Citované 2024-05-12] Dostupné na https://matplotlib.org/.
- Brian Meloon and Julien C Sprott. Quantification of determinism in music using
iterated function systems. Empirical Studies of the Arts, 15(1):3–13, 1997.
- Qingxi Meng, Shubham Chandak, Yifan Zhu, and Tsachy Weissman. Reference-free lossless compression of nanopore sequencing reads using an approximate assembly approach. Scientific Reports, 13(1):2082, 2023.
- Knižnica numpy. [Citované 2024-05-12] Dostupné na https://numpy.org/.
- National Institutes of Health et al. International nucleotide sequence database
collaboration. [Citované 2024-04-22] Dostupné na https://www.insdc.org/.
- David F Robinson and Leslie R Foulds. Comparison of phylogenetic trees. Mathematical biosciences, 53(1-2):131–147, 1981.
- Jill Roughan. Your essential guide to different file formats in bioinformatics. [Citované 2023-05-27] Dostupné na https://www.formbio.com/blog/your-essential-guide-different-file-formats-bioinformatics.
- Hiruna Samarakoon, Hasindu Gamaarachchi, et al. Flexible and efficient handling
of nanopore sequencing signal data with slow5tools. Genome Biology, 24(1):69,
2023.
- Muhammad Sardaraz and Muhammad Tahir. SCA-NGS: Secure compression algorithm for next generation sequencing data using genetic operators and block
sorting. Science Progress, 104(2):00368504211023276, 2021.
- Knižnica scipy. [Citované 2024-05-12] Dostupné na https://scipy.org/.
- Milton Silva, Diogo Pratas, and Armando J Pinho. Efficient DNA sequence compression with neural networks. GigaScience, 9(11):giaa119, 2020.
- Shannon M Soucy, Jinling Huang, and Johann Peter Gogarten. Horizontal gene transfer: building the web of life. Nature Reviews Genetics, 16(8):472–482, 2015.
- Catalin Stoean and Daniel Lichtblau. Author identification using chaos game
representation and deep learning. Mathematics, 8(11):1933, 2020.
- Susana Vinga. Alignment-free methods in computational biology, 2014.
- Susana Vinga and Jonas Almeida. Alignment-free sequence comparison—a review.
Bioinformatics, 19(4):513–523, 2003.
- Aimin Yang, Wei Zhang, Jiahao Wang, Ke Yang, Yang Han, and Limin Zhang.
Review on the application of machine learning algorithms in the sequence data
mining of dna. Frontiers in Bioengineering and Biotechnology, 8:1032, 2020.
- Andrzej Zielezinski et al. Afproject. [Citované 2024-03-19] Dostupné na https://afproject.org/app/.
- Andrzej Zielezinski et al. Benchmarking of alignment-free sequence comparison
methods. Genome biology, 20(1):1–18, 2019.
- Andrzej Zielezinski, Susana Vinga, Jonas Almeida, and Wojciech M Karlowski.
Alignment-free sequence comparison: benefits, applications, and tools. Genome
biology, 18:1–17, 2017.